Transcriptome-Derived SSR Marker Development for Genetic Insights into Dipterocarpus alatus Roxb. ex G.Don, a Vulnerable and Ecologically Significant Species
DOI:
https://doi.org/10.4238/37e0sz96Keywords:
Dipterocarpus alatus, genic SSR markers, Transcriptome sequencing, Genetic diversity, Population structure, Vulnerable species, Conservation geneticsAbstract
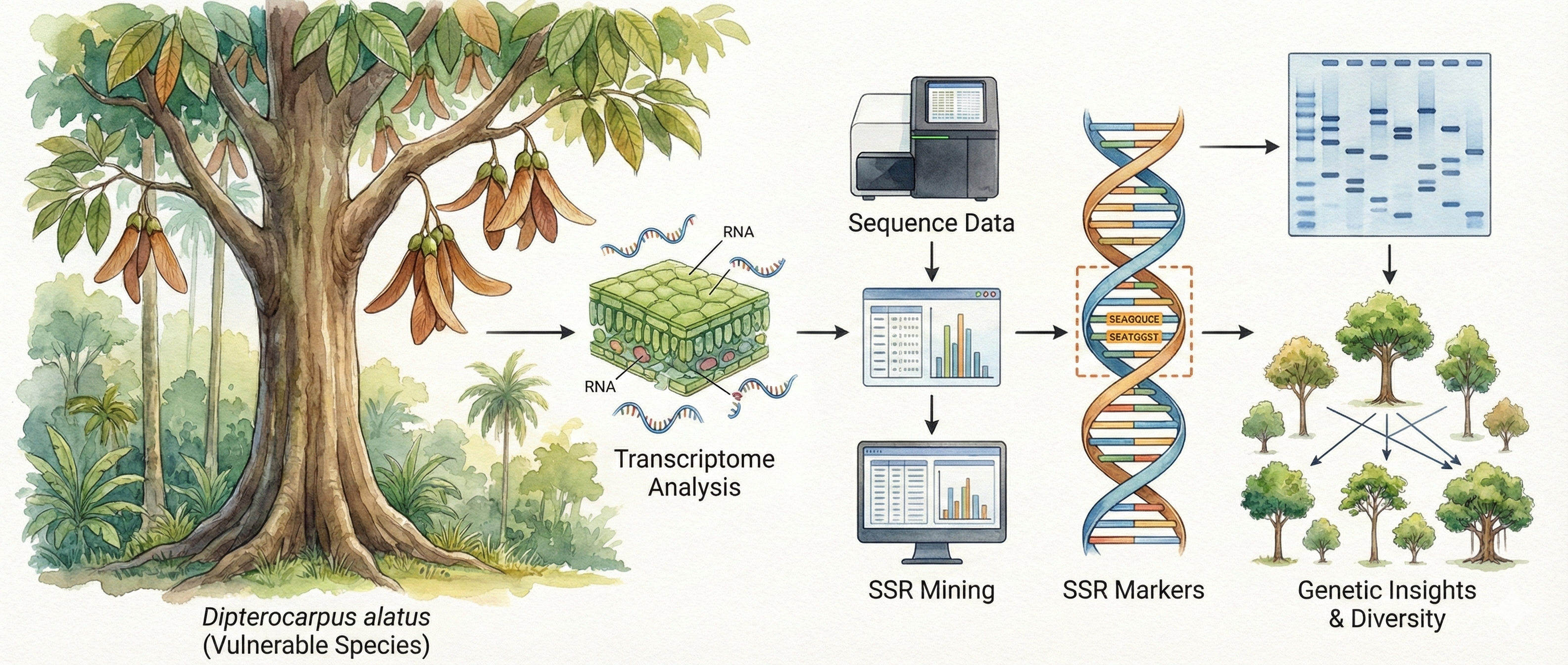
Dipterocarpus alatus Roxb. ex G.Don is a tropical tree species of significant ecological and economic importance that is increasingly threatened by habitat fragmentation and overexploitation. However, there are no efficient tools to estimate the population structure and genetic diversity of D. alatus. Here, we employed a simple sequence repeat (SSR) marker development method derived from the leaf transcriptome sequences and validated for the analysis of genetic diversity and population structure. Over 48 million clean reads were generated from the transcriptome of the young leaf tissue. A total of 12,514 SSR loci were identified from the assembled transcripts. Among them, mononucleotide repeats were most abundant (75.32%), followed by trinucleotide (15.76%), dinucleotide (7.89%), tetranucleotide (0.81%), hexanucleotide (0.13%), and pentanucleotide (0.09%) motifs. Although mononucleotide repeats were predominant, only di-, tri, and tetranucleotide motifs were selected for marker development due to their higher suitability. 61 loci were chosen to screen the polymorphic information, and 26 of them yielded reproducible amplification across 30 individuals from five D. alatus populations. These markers exhibited polymorphism information content (PIC) values ranging from 0.315 to 0.410, with an average discriminatory power (Dp = 0.385), indicating that these markers are informative and suitable for assessing genetic variation among individuals. Trinucleotide repeats, often located in coding regions, demonstrated high success rates in primer design. Genetic diversity analysis revealed observed heterozygosity (Ho) ranging from 0.000 to 0.823 and expected heterozygosity (He) from 0.036 to 0.770. Population structure analysis revealed three genetically distinct clusters that largely corresponded to geographic origins. The presence of admixed individuals and within-cluster substructuring suggests ongoing gene flow and complex genetic relationships among populations. This study presents the first set of transcriptome-based genic SSR markers for D. alatus, offering valuable molecular tools for conservation genetics and sustainable forest resource management.
Downloads
Published
Issue
Section
License
Copyright (c) 2025 Preeya Wangsomnuk, Xin Chun Mo, Manop Poopath (Author)

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.



